An extended SEIQR type model is considered in order to model the COVID-19 epidemic. It contains the classes of susceptible individuals, exposed, infected symptomatic and asymptomatic, quarantined, hospitalized and recovered. The basic reproduction number and the final size of epidemic are determined. The model is used to fit available data for some European countries. A more detailed model with two different subclasses of susceptible individuals is introduced in order to study the influence of social interaction on the disease progression. The coefficient of social interaction K characterizes the level of social contacts in comparison with complete lockdown (K=0) and the absence of lockdown (K=1). The fitting of data shows that the actual level of this coefficient in some European countries is about 0.1, characterizing a slow disease progression. A slight increase of this value in the autumn can lead to a strong epidemic burst.
1.
Introduction
The Coronavirus disease 2019 (COVID-19) pandemic is now considered as the biggest global threat worldwide. This disease is caused by severe acute respiratory syndrome coronavirus 2 (SARS-COV-2) which belongs to a group of RNA virus causing respiratory track infection that can range mild to lethal [1]. The first outbreak of COVID-19 was noticed in Hubei province, Wuhan, China [2] in December 2019. Then it spread all over the world so rapidly that the World Health Organization (WHO) revealed the COVID-19 to be a public health emergency and identified it as a pandemic on March 11, 2020. Since December 2019, the first COVID-19 infected person was diagnosed, the COVID-19 quickly spread to all Chinese province and, as of 1 April 2020, to 200 countries and regions with the number of reported infected cases and the number of documented death reached 19 million and 717, 792, respectively [3]. Among these countries, the alarming epidemic situations are in the United States (5, 032, 278 total cases, 162, 804 death cases), Brazil (2, 917, 562 total cases, 98, 644 death cases), India (2, 030, 001 total cases, 41, 673 death cases), Russia (871, 894 total cases, 14, 606 death cases), Italy (249, 204 total cases, 35, 187 death cases), UK (308, 134 total cases, 46, 413 death cases), Spain (354, 530 total cases, 28, 500 death cases), Germany (215, 210 total cases, 9, 252 death cases), France (195, 633 total cases, 30, 312 death cases), and Iran (320, 117 total cases, 17, 976 death cases) as of 8th August, 2020. In particular, the number of infection cases in the United States has grown very fast, the number of reported infected cases increases from 15 to 288, 721 spending 82 days only [3].
The most alarming part of this disease spread is that the symptoms of this disease are not specific and in many cases the infected person may be asymptomatic (who can infect other persons without showing any symptoms of the disease). The majority of the cases initially have symptoms like common cold which includes dry cough, fever, sore throat, loss of sense of smell, headache, shortness of breathe etc. Moreover, the disease growth of this infection further proceed to acute respiratory distress syndrome which can lead to even death. It is also observed in an age-stratified analysis [4] that the large number of severe cases in particular for the age groups above 60 and having other medical issues like diabetics, kidney problems etc. The COVID-19 virus spreads at large extend between people when they come in close contact with each other and the virus is transmitted through expelled droplets which enter a person's body through various contact routes such as the mouth, eyes or nose. Contact with various surface is another means for contracting the virus. In the absence of a definite treatment modality like appropriate/effective medicine or vaccine, physical distancing, wearing masks, washing hands etc. have been accepted globally as the most efficient strategies for reducing the severity and spread of this virus and gaining control over it at some extend [5]. So, the governments of most of the countries had decided to go for complete lockdown from mid or end of March, 2020. This complete lockdown also affected the economy of those countries in a large extent. So, from the mid of the month of May, 2020, all the countries have taken some policies for unlocking gradually.
Mathematical models of infectious disease dynamics nowadays became a very useful and important tool for the analysis of dynamics of disease progression, to predict the future course of an outbreak and to evaluate strategies to control an epidemic in recent years. The global problem of the outbreak of COVID-19 has attracted the interest of researchers of different areas. Mathematical modeling based on system of differential equations may provide a simple but comprehensive mechanism for the dynamics of COVID-19 transmission. Several modeling studies have already been performed for the COVID-19 outbreak [6,7,8,9,10,11]. In [12], Lin et. al. suggested a conceptual model for the coronavirus disease started at the end of 2019, which effectively catches the time line of the COVID-19 outbreak. A mathematical model for reproducing the stage-based transmissibility of a novel coronavirus is proposed and analyzed by Chen et al. in [13]. Wu et al. developed a susceptible exposed infectious recovered model (SEIR) based on the reported data from December 31st, 2019 till January 28th, 2020, to clarify the transmission dynamics and projected national and global spread of the disease [14]. They also calculated the basic reproduction number as around 2.68 for COVID-19. Tang et al proposed a compartmental deterministic model that would combine the clinical development of the disease, the epidemiological status of the patient and the measures for intervention. Researchers also found that the value of control reproduction number may be as high as 6.47, and that methods of intervention including intensive contact tracing followed by quarantine and isolation would effectively minimize COVID-19 cases [15]. For the basic reproductive number (R0), Read et al. reported a value of 3.1 based on the data fitting of an SEIR model, using an assumption of Poisson-distributed daily time increments [16]. S. Zhao et al. estimated the mean R0 for 2019-nCoV in the early phase of the outbreak ranging from 3.3 to 5.5 (likely to be below 5 but above 3 with rising rate of reported cases) [17], which appeared slightly higher than those of SARS-CoV (R0: 2–5) [18]. A report by Cambridge University has indicated that India's countrywide three-week lockdown would not be adequate to prevent a resurgence of the new coronavirus epidemic that could bounce back in months and cause several thousands of infections [19]. They suggested that two or three lockdowns can extend the slowdown longer with five-day breaks in between or a single 49-day lockdown. Data-driven mathematical modeling plays a key role in disease prevention, planning for future outbreaks and determining the effectiveness of control. Several data-driven modeling experiments have been performed in various regions [15]. In [20], a compartmental mathematical model is developed to understand the outbreak of COVID-19 in Mexico. This data driven analysis would let us compare how different the outbreak will be in the two studied regions. By this approach, authorities can plan a health care program to control the spread even with limited resources. In Rojas et al. [21], the authors estimated the value of R0 which helped them to predict that in the city of Cali the outbreak under current intervention of isolation and quarantine will last for 5–6 months and will need around 3500 beds on a given during the peak of the outbreak. Some other relevant works where the basic reproduction number is estimated for different countries can be found in [22,23,24,26,25,27,28].
The most common mathematical formulations which represent the individual transition in a community between 'compartments' describe the situation of individual infection with a significant insight. These compartmental models for disease progression segregate a population into groups depending on each individual's infectious state and related population sizes with respect to time. There is a wide range of mathematical models and approaches adopted by different researchers to develop viable mathematical model to understand the propagation of disease spread for COVID-19 with different model assumptions. In our present work we have proposed and analyzed an ordinary differential equation model to study the COVID-19 disease propagation which consists with initially six compartments namely susceptible, exposed, infected, quarantined, hospitalised and recovered. We have divided the exposed compartment into two sub-compartments (denoted by E1 and E2) depending on their infectiousness and also divided the infected class into two sub-compartments, namely, asymptomatic (Ia) and symptomatic (Is). Interesting and significant contribution of our work is the consideration of time dependent rate of infection over various periods of time. This variation is adopted into the model in order to capture the effect of lockdown, social distancing etc. which plays a crucial role to reduce the disease spread. Next we have discussed the basic properties of our model and calculated the basic and controlled reproduction number. Final size of the epidemic is also described. We have divided susceptible, exposed and infected classes into two sub-classes each based on their classification or behaviour which is directly responsible for the alteration of rate of disease spread. The classification or division into two different groups may be due to the different age groups, different implementation of distancing measures, proper and improper use of face mask and so on. Then we have calculated the reproduction number and final size of the epidemic for the two group model. Next we have copulated the sensitivity index to identify the parameters of greater interest and then fitted those parameter values with the data of total cumulative cases of COVID-19 for different countries (Germany, Italy, Spain, and UK). Then we have shown that with those best fitted parameter values our model simulation is matching well with the 95% confidence interval of the daywise cumulative and daily data for reported cases which proved the viability of our model system. This also depict the fact that nature of the disease transmission is different for different countries depending on the protective measures and policies taken by government, different age distribution of the population, lack of consciousness and many others.
2.
Mathematical model – 1
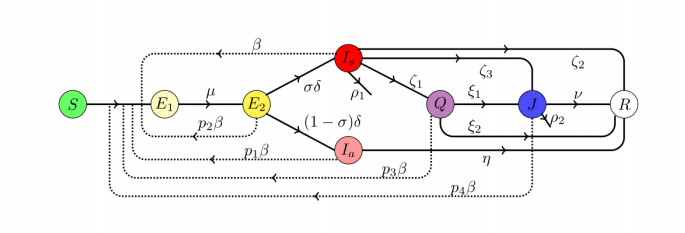
In the following, we consider a dynamic SEIQR type model for the COVID-19 disease progression. Basically the model consists with Susceptible (S), Exposed (E), Infected (I), Quarantined (Q), hospitalized (J) and Recovered (R) class. In the context of COVID-19, the exposed class is divided into two subclasses namely non-infectious (E1) and infectious (E2) and the infected class is divided into two sub- classes namely asymptomatic (Ia) and symptomatic (Is), where N=S+E1+E2+Ia+Is+Q+J+R and N is not fixed since deceased individuals are not considered in the model explicitly. The model assumptions are given as follows:
Susceptible population S(t): This subpopulation will decrease after an infection due to the interaction with an symptomatic infected individual (Is), asymptomatic infected individual (Ia), infectious exposed individual (E2), quarantined (Q) or hospitalised one (J). The transmission coefficients will be βIs/N, βp1Ia/N, βp2E2/N, βp3Q/N and βp4J/N respectively. Here β is rate of infection per unit of time by the symptomatic infected, p1, p2, p3 and p4 are the reduction factor of infectivity by Ia, E2, Q and J respectively compared to Is and satisfy the restriction 0≤pj<1, j=1,2,3,4. The rate of change of the susceptible population is expressed in the following equation:
Exposed polulation E(t): The exposed population (in the incubation period) is divided into two sub classes: (i) exposed population who are at the beginning of the incubation period and cannot spread the disease (E1(t)) and (ii) exposed population who are at the end of the incubation period and can spread the disease (E2(t)), hence E(t)=E1(t)+E2(t). The transfer mechanism from the class S(t) to the class E1(t) is guided by the function βSN(Is+p1Ia+p2E2+p3Q+p4J) and from the class E1(t) to the class E2(t) is guided by μE1, where μ is the rate at which individuals of E1 class become infectious exposed (E2). The population of E2 will decrease due to the transfer into the infected class with rate δ. Thus the rate of change of the exposed population is expressed in the following two equations:
Infected polulation I(t): The infected population is divided into two subclasses (i) asymptomatic (having no symptoms) (Ia(t)) and (ii) symptomatic (having symptoms) (Is(t)), such that I(t)=Ia(t)+Is(t). The transfer mechanism from the class E2(t) to the class Ia(t) is guided by the function (1−σ)δE2 and from the class E2(t) to the class Is(t) is guided by the function σδE2 where σ (0<σ<1) is the fraction of E2 that becomes symptomatic infected (Is). The population Ia will decrease due to the transfer into the recovered population with a rate η and Is will decrease with a rate (ρ1+ζ1+ζ2+ζ3), where ρ1 is the rate of mortality due to the infection, ζ1, ζ2 and ζ3 are the rate at which Is becomes quarantined, recovered and hospitalised respectively. Thus the rate of change of the infected population is expressed in the following two equations:
Quarantined population Q(t): This subpopulation will increase due to the transfer from the class Is(t) with a rate ζ1 and will decrease with a rate (ξ1+ξ2), where ξ1 and ξ2 are the rates at which Q becomes hospitalized and recovered respectively. Thus the rate of change of the quarantined population is expressed in the following equation:
Hospitalized population J(t): This subpopulation will increase due to the transfer from the classes Is(t) and Q(t) with rates ζ3 and ξ1 respectively and will decrease with a rate (ρ2+ν), where ρ2 is the rate of mortality due to the infection and ν is the rate at which the hospitalized individuals are recovered. Thus the rate of change of the hospitalized population is expressed in the following equation:
Recovered population R(t): This subpopulation will increase due to recovery from the disease from the classes Ia(t), Is(t), Q(t) and J(t) with rates η, ζ2, ξ2 and ν respectively. Thus the rate of change of the recovered population is expressed in the following equation:
Hence, the system of differential equations that will model the dynamics of coronavirus spread is:
subjected to non-negative initial conditions S(0),E1(0),E2(0),Ia(0),Is(0),Q(0),J(0),R(0)≥0.
Interpretation for the parameters involved with the model (2.1) is summarized in Table 1 for a quick reference. A schematic diagram for the transmission of disease and progression of the individuals from one compartment to another is provided in Figure 1.
We have written the basic model with β as constant for the simplicity of forthcoming mathematical calculations. However, for numerical simulations and in order to fit the numerical results with available data [3], we will consider β≡β(t) as a function of time in order to model the effect of lockdown. In reality the rate of infection is not a constant throughout the epidemic outbreak rather it changes time to time due to the variability in social behavior. Although it is difficult to obtain actual pattern of variation with respect to time but we have considered this variation in order to model the lowering in rate to infection due to the lowering of social contancts during the lockdown period.
2.1. Positivity and boundedness
A viable mathematical model for epidemiology must ensure that the solutions of the model under consideration remain non-negative once started from an interior point of the positive cone and remains bounded at all future time. The model considered here is not a completely new rather several close versions are available in various articles on mathematical epidemiology. However, for the completeness we just state the relevant result and a brief outline for the proof is provided at the appendix. For this purpose we consider that the model (2.1) is subjected to the initial conditions (S(0),E1(0),E2(0),Ia(0),Is(0),Q(0),J(0),R(0))∈R8+, where positive cone of R8 is R8+={(x1,...,x8):xi≥0,i=1,...,8}. First we prove that the solution of the system (2.1) remains within R8+ at all future time once started from a point within R8+ following the approach outlined in [29].
Theorem 1. The system (2.1) is invariant in R8+.
Proof. See appendix.
In order to close the model (2.1), we can introduce the compartment for number of COVID related deaths as follows
Now if we write ND=S+E1+E2+Ia+Is+Q+J+R+D, then from (2.1) and (2.1d), we find
With the previous urgument we can prove D(t)≥0 and hence 0<N≤ND implies N is bounded above, consequently all the constituent variables are also bounded.
2.2. Basic and controlled reproduction numbers
In this section, first we find the basic reproduction number for the model (2.1) in the absence of any control measures, namely, quarantine and hospitalization. For this purpose we assume that the model consists with Susceptible, Exposed, Infected and Recovered class only. The Quarantined and Hospitalized classes are not considered for the time being. Based upon this assumption we can write the following reduced model:
subjected to non-negative initial conditions S(0),E1(0),E2(0),Ia(0),Is(0),R(0)≥0. For this model N=S+E1+E2+Ia+Is+R. Here we calculate the basic reproduction number for above model following the next generation matrix approached as introduced in [32]. The disease free equilibrium point is given by (N,0,0,0,0,0), the derivation of basic reproduction number is based upon the stability condition of disease free equilibrium point for the model (2.4). First we rearrange the compartments of the model (2.4) such that first four equations are related to infected compartments from epidemiological point of view (i.e., E1, E2, Ia and Is). Susceptible and recovered compartments will be placed at the end. From (2.4), we can write
where X=[E1,E2,Ia,Is,S,R]T≡[x1,x2,x3,x4,x5,x6]T. Here the matrix F1 consists of the terms involved with the appearence of new infection at all the compartments and V1 contains the terms representing entry and exit of all other individuals, rather than direct infection, at all compartments. Explicit expressions for F1 and V1 are given at the appendix.
For the calculation of basic reproduction number now we need to evaluate two matrices F1 and V1, which are defined by
where X0=(0,0,0,0,N,0)T. Here we should mention that F1≡[f11,f12,f13,f14,f15,f16]T and V1≡[v11,v12,v13,v14,v15,v16]T to avoid any confusion. The largest eigenvalue of the next generation matrix F1V−11 is the basic reproduction number for the model (2.4), (see [32,33]). The matrices F1, V1 and F1V−11 are given at the appendix.
The basic reproduction number for the model (2.4) is denoted by R[1]0 and is given by
Here the superscript ' [1] ' stands for the first model considered in this manuscript, that is for the model (2.1). This basic reproduction number is the sum of three terms, which represents the number of secondary infections produced by an infectious exposed individual, an asymptomatic infected individual, and a symptomatic infected individual respectively. βp2SE2N is the incidence of an exposed individuals who are at the end of the incubation period and can spread the disease. The number of secondary infection produced by an individual of E2 compartment in an entirely susceptible population is βp2 per unit of time. An individual spents an average 1δ units of time in E2 compartment. Hence the number of secondary infection produced by an individual of E2 compartment is βp2δ. The number of secondary infections produced by the individuals of Ia and Is compartments in an entirely susceptible population, per unit of time, are β(1−σ)p1 and βσ respectively. The average time units spend by the asymptomatic and symptomatic infectious individuals with their respective compartments are 1η and 1ρ1+ζ2. Hence the number of secondary infection produced by the individuals of Ia and Is comparments are β(1−σ)p1η and βσρ1+ζ2 respectively.
Henceforth we follow the similar notation and approach to calculate other relevant reproduction numbers without providing detailed description, the expressions for large matrices are provided at the appendix. Now we calculate the controlled reproduction number for the model (2.1).
The model (2.1) can be written as follows
X=[E1,E2,Ia,Is,Q,J,S,R]T≡[x1,x2,x3,x4,x5,x6,x7,x8]T with six compartments contribute to the propagation of infection. We can define following two matrices
where X0=(0,0,0,0,0,0,N,0).
The controlled reproduction number for the model (2.1) is given by
The different terms involved with R[1]c can be explained in a similar way as given above, interested readers can see [34,35] for detailed discussion. It is important to mention here that substituting ζ1=ζ3=0 we can find the basic reproduction number R[1]0 from the controlled reproduction number R[1]c.
2.3. Final sizes of epidemic
In order to determine the final size of the epidemic, we find three relevant quantites Sf, Rf and Df. Here Sf is the final size of the susceptible compartment at the end of outbreak, theoretically it is defined by limt→∞S(t)=Sf [34,35]. Let tf denotes the time at which the number of infected is zero that is at the end of epidemic outbreak and consequently Sf, Rf and Df can be considered as the values of S(t), R(t) and D(t) at t=tf. First we calculate the final size of the susceptible compartment that is Sf. We integrate the equation for S (i.e., (2.1a)) between t=0 to t=tf(=∞) and find
where S0 and Sf denotes initial and final size of the susceptible population.
Now we assume that the model (2.1) is subjected to the initial condition that S0,E10>0 and all other components are absent at the initial time point t=0. Consequently S0+E10=N. Now integrating (2.1c) between t=0 and t=tf and with the assumption that E20=E2f=0, we find
Similarly from (2.1d) we find the following result,
Proceeding in a similar way, from Eqs (2.1e)–(2.1g) and using above results we find,
and
Now if we add two equations (2.1a) and (2.1b), and then integrating we find
where S0=N and we assume E10,E1f=0. Now using the results (2.11)–(2.15), from (2.10) we find
Finally using (2.16) and the expression for R[1]c, we obtain the final size of the epidemic as follows
At the begining of the epidemic without any loss of generality we can assume that the entire population is susceptible and hence N=S0. Using SfS0≡SfN=x in above equation we get the following equation
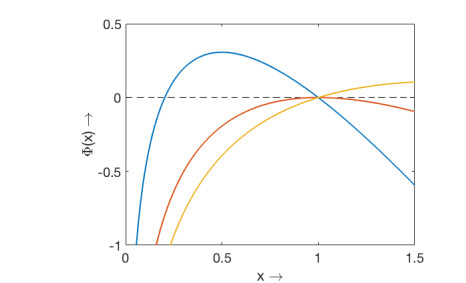
This equation possesses a solution within the interval (0,1) when R[1]c>1 and root of the equation Φ(x)=0 gives the final size of the epidemic. On the other hand the question of final size of the epidemic will not arise in case of R[1]c≤1 as there is no root of the equation Φ(x)=0 within (0,1). This claim can be verified from the Figure 2.
To understand the final size of the deceased compartment we need to calculate Df and Rf. Clearly R0=0 and D0=0, hence by integrating the equations for recovered and deceased compartments between t=0 to t=tf we find
and
Using the results (2.12)–(2.15), we find from above two equations,
and
If we define
then
3.
Mathematical model – 2
In the following, we have divided the susceptible population (S), Exposed population (cannot spread infection (E1) and can spread infection (E2)), asymptomatic infective (Ia) and symptomatic infective (Is) of model (2.1) into two subclasses, namely S1, S2, E11, E21, E12, E22, Ia1, Ia2, Is1, Is2 respectively based on their classification or behaviour which is directly responsible for the alteration of rate of disease spread. The classification or division into two different groups may be due to the different age groups, different implementation of distancing measures, proper and improper use of face mask and so on, related to the human behaviours. The total population size can be written as N=S1+S2+E11+E21+E12+E22+Ia1+Ia2+Is1+Is2+Q+J+R and N is not fixed since deceased compartment is not included in the model. The assumptions for the extended model formulation are given as follows:
Susceptible population S(t): This subpopulation is divided into two subclasses S1 and S2. The population in the compartment S1 will decrease after an infection due to the interaction with an symptomatic infected individual (Is1,Is2) with transmission coefficients β11Is1/N, β12Is2/N respectively, asymptomatic infected individual (Ia1,Ia2) with transmission coefficients β11p11Ia1/N, β12p21Ia2/N respectively, infectious exposed individual (E12,E22) with transmission coefficients β11p12E12/N, β12p22E22/N respectively, quarantine (Q) with transmission coefficients βQQ/N or hospitalised one (J) with transmission coefficients βJJ/N. Here β11, β12 are rates of infection per unit of time by the symptomatic infected Is1, Is2 respectively, βQ, βJ are rates of infection per unit of time by the quarantined and hospitalized population (Q,J) respectively, p11, p12, p21 and p22 are the reduction factor of infectivity by Ia1, E12, Ia2 and E22 respectively compared to Is1 and Is2 and satisfy the restriction 0≤pij<1, i,j=1,2. S2 will decrease after an infection due to the interaction with an symptomatic infected individual (Is1,Is2) with transmission coefficients β21Is1/N, β22Is2/N respectively, asymptomatic infected individual (Ia1,Ia2) with transmission coefficients β21p31Ia1/N, β22p41Ia2/N respectively, infectious exposed individual (E12,E22) with transmission coefficients β21p32E12/N, β22p42E22/N respectively, quarantine (Q) with transmission coefficients βQQ/N or hospitalised one (J) with transmission coefficients βJJ/N. Here β21, β22 are rates of infection per unit of time by the symptomatic infected Is1, Is2 respectively, βQ, βJ are rates of infection per unit of time by the quarantined and hospitalized population (Q,J) respectively, p31, p32, p41 and p42 are the reduction factor of infectivity by Ia1, E12, Ia2 and E22 respectively compared to Is1 and Is2 and satisfy the restriction 0≤pij<1, i=3,4,j=1,2. The rate of change of the susceptible population is expressed in the following two equations:
The rate of change of two groups of the exposed compartments, who are not infectious, is described by the follwoing two equations
where μ1 and μ2 are the rates at which E11 and E21 progress to the infectious exposed compartments E12 and E22 respectively. The infectious exposed individuals leave the respective classes at rates δ1 and δ2, hence their rate of change with respect to time is given by
The governing equations for the two groups of asymptomatic and symptomatic infected classes are given as follows:
where the parameters σj, ρj1, ζj1, ζj2 and ζj3 (j=1,2) have the similar interpretation as that of σ, ρ1, ζ1, ζ2 and ζ3 as described in Table 1 for model (2.1).
Distinction for quarantined, hospitalized and recovered compartments are not required as first two groups are under indirect and direct medical interventions and recovered individuals have no direct role to play with the disease spread. Finally including the governing equations for quarantined, hospitalized and recovered compartments and using previous equations we get the final two-group infection model as follows,
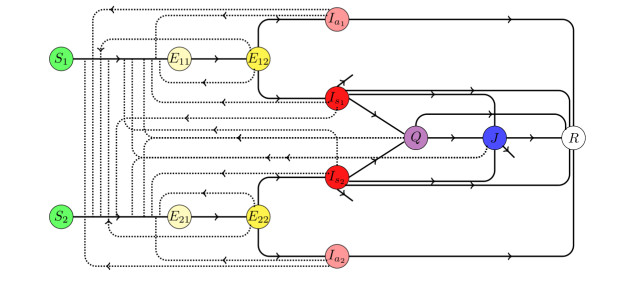
Here ρ11, ρ21, ξ1 and ρ2 are the disease related death rates for the compartments Is1, Is2, Q and J respectively. A schematic diagram is presented in Figure 3 without the rate constants in order to avoid clumsiness.
Before analyzing the above model, here we explain how the model (2.1) can be derived from the model (3.1) under suitable assumptions. For this purpose first adding the Eqs (3.1e) and (3.1f), we find
If we assume μ1=μ2=μ and δ1=δ2=δ, then we find
which is Eq (2.1c) if we write (E11+E21)=E1 and (E12+E22)=E2 respectively.
Similarly, with the previous assumptions on parameters and variables, and additional assumptions σ1=σ2=σ, η1=η2=η and Ia1+Ia2=Ia, we can derive (2.1d) by adding (3.1g) and (3.1h). We can also derive (2.1e) from the Eqs (3.1i) and (3.1j) using the similar procedure.
Derivation of Eq (2.1a) by adding Eqs (3.1a) and (3.1b) is little bit tricky. Adding (3.1a) and (3.1b), and rearranging the terms, we can write
Firstly, if we assume β11=β12=β21=β22=β, βQ=p3β, βJ=p4β, then we can write
Further if we assume pj1=p1, pj2=p2, j=1,2,3,4, we can write from above equation
Finally writing S1+S2=S, and using previous assumptions related to E2, Ia and Is, we can write above equation as
With the other necessary assumptions on the remaining parameters and variables we can derive the model (2.1) from (3.1). Once the two group of individiuals are non-distinguishable, then the model (2.1) results in from the model (3.1).
3.1. Controlled reproduction number
For simplicity of forthcoming calculations, we can write the first two Eqs (3.1a)–(3.1b) into the matrix form as follows:
where
and
Using above approach, we can write (3.1c)– (3.1d) into a compact form as follows
where
Similarly, Eqs (3.1e)–(3.1f), (3.1g)–(3.1h) and (3.1i)–(3.1j) can be written as
and
respectively.
In order to calculate the controlled reproduction number for the model (3.1), we follow the similar approach as we have used for the model (2.1). Without giving much description, here we just define the relevant matrices as follows,
Evaluating the Jacobian matrices, at (N1,N2,0,0,0,0,0,0,0,0,0,0,0) (the disease free equilibrium point), corresponding to F3 and V3, we can find F3 and V3, their explicit expressions are provided at the appendix. The controlled reproduction number is the largest eigenvalue of the matrix F3V−13.
In terms of the matrices introduced above, we can rewrite F3 and V3 as partitioned matrices analogous to the matrices F2 and V2 as in the previous section. F3 and V3 can be rewritten as
where
Two constants α1 and α2 are defined by
The matrix V−13 is a lower triangular partitioned matrix. Based upon the non-zero entries of F3, we need the elements in first two columns of the matrix V−13 in order to calculate the controlled reproduction number. If we write V−13 in the following form
where
Once we calculate the matrix F3V−13, it comes out to be a matrix of following form
and hence the non-zero eigenvalues can be determined from the first 2×2 block of the block matrix Γ2×10 involved with F3V−13. The entries of the first 2×2 block can be calculated as follows,
Explicit expressions for γij, (i,j=1,2) are given by
The matrix F3V−13 has at most two non-zero eigenvalues and eight zero eigenvalues. The maximum positive eigenvalue of the matrix F3V−13 is the controlled reproduction number for the model (3.1). As a matter of fact the controlled reproduction number for the model (3.1) is the largest eigenvalue of the matrix [γij]2×2 and is given by
Here the superscript '[2]' stands for the second mathematical model considered in this manuscript, that is for the model (3.1).
3.2. Final size of epidemic
Derivation of final size of the epidemic is tedious but can be obtained through step by step calculations. The final sizes, for different compartments, can be determined in a similar manner as we have done in the previous section. From (3.1a) & (3.1b), integrating between the limits t=0 and t=tf(=∞), we find
where Sj0 and Sjf denote initial and final size of the j-th susceptible class, j=1,2. Now we assume that the model (3.1) is subjected to the initial conditions such that S10,S20,E110,E210>0 and all other components are absent at the initial time point t=0. Consequently S10+S20+E110+E210=N. Now integrating (3.1e), (3.1f) between t=0 and t=tf and with the assumption that E120=E220=E12f=E22f=0, we find
Next integrating (3.1g), (3.1h) between t=0 and t=tf and with the assumption that Ia10=Ia20=Ia1f=Ia2f=0, using (3.25) we find
Proceeding in a similar way and using the results in (3.26), we get from (3.1i), (3.1j)
where α1 and α2 are defined in (3.16). Using the above result, from (3.1k) we get
Finally, integrating (3.1l) between t=0 and t=tf and using J0=0=Jf, we can write
Using the results from (3.27) and (3.28), we can express the ∫tf0Jdt in terms of ∫tf0E11dt and ∫tf0E21dt. Now adding the equations (3.1a) and (3.1c) and then integrating between t=0 and t=tf, we find
where we have used E110=E11f=0 and S10=N1. Similarly, adding the Eqs (3.1b) and (3.1d) and following the same approach, we can find
Now using the results (3.25)–(3.29), we get from (3.23),
where
Proceeding in a similar way, from (3.24), we can derive
where
Eliminating ∫tf0E11dt and ∫tf0E21dt between (3.30), (3.31) and (3.35), we find the following two equations
Without any loss of generality, writing S10=N1 and S20=N2 and introducing the notations S1fN1=x, S2fN2=y we get from above system equations
where
It is interesting to note that the following two matrices are similar matrices,
If we define a diagonal matrix
then we can verify that
Entries of both the matrices P and Q are positive and as they are similar, the spectrum of two matrices are same. Hence the largest eigenvalue of the matrix Q is equal to the largest eigenvalue of the matrix P that is equal to R[2]c.
The existence of final size of the epidemic depends upon the existence of a point of intersection between two curves y=Φ1(x) and x=Φ2(y) within the unit square [0,1]×[0,1]. Existence of such point of intersection follows from the Th. 1 in [36]. The two curves y=Φ1(x) and x=Φ2(y) intersect each other at a point (ˉx,ˉy), 0<ˉx,ˉy<1 whenever R[2]c>1. Knowing the values of ˉx, ˉy we can determine the final sizes S1f and S2f.
4.
Numerical simulations
In this section we present the numerical simulation results for two models (2.1) and (3.1) considered in this work. First we present the simulation results for the model (2.1) by fitting the numerical simulation results with the COVID-19 data for Germany as an illustrative example. All the data used in this manuscript are taken from [3]. In order to ensure that the obtained result and the validity of the model is not country specific rather it can be matched with the data for other countries. Fitting of the data for Italy, UK and Spain are provided at the appendix.
The sensitivity of various parameters involved with the model (2.1) plays an important role behind numerical simulations. It would be quite lengthy calculations if we want to perform the same for the model (3.1) and hence we have restricted ourselves to the first model only. For numerical simulations and fitting with the data, we have considered the rate of infection as a function of time and hence some of the estimated parameters do not remain fixed throughout the simulation and hence we have presented the sensivity indices with respect to the estimated parameter values.
4.1. Sensitivity analysis
The sensitivity analysis for the endemic threshold (the controlled reproduction number R[1]c) in Eq (2.9) tells us how important each parameter is to disease transmission. It is used to understand parameters that have a high impact on the threshold R[1]c and should be targeted by intervention strategies. More precisely, sensitivity indices’ allows us to measure the relative change in a variable when a parameter changes. For that purpose, we use the normalized forward sensitivity index of a variable with respect to a given parameter, which is defined as the ratio of the relative change in the variable to the relative change in the parameter. If such variable is differentiable with respect to the parameter, then the sensitivity index is defined as follows.
The normalized forward sensitivity index of R[1]c, which is differentiable with respect to a given parameter θ (say), is defined by
From direct calculation and without using the numerical values we can determine the signs of sensitivity indices with respect to some of the parameters. For the remaining sensitivity indices, we need to take help of the numerical examples. The normalized forward sensitivity indices of R[1]c with respect to the parameters which are found to be clearly positive are given by,
where A1=ρ1+ζ1+ζ2+ζ3, A2=ξ1+ξ2 and A3=ρ2+ν.
The normalized forward sensitivity indices of R[1]c with respect to the parameters which are found to be clearly negative are given by,
The signs of normalized forward sensitivity indices of R[1]c with respect to rest of the parameters can not be determined from their explicit expressions and we need to take help of some numerical examples. Such normalized forward sensitivity indices are as follows,
We use the sensitivity indices to understand parameters that have a high impact on R[1]c. The values of the sensitivity indices for different parameters depend on the choice of parameter values. As we have mentioned earlier, the values of the sensitivity indices can be positive or negative. The most positive (or negative) value of the sensitivity index for a parameter indicates that parameter is most sensitive to R[1]c and the least positive (or negative) value of the sensitivity index for a parameter indicates that parameter is least sensitive to R[1]c.
The sensitivity index may depend on several parameters of the system, but also can be constant, independent of any parameter. For example, ΥR[1]cβ=1 means that an increase (decrease) in β by a given percentage will result in increase (decrease) in R[1]c by the same percentage. The estimation of a sensitive parameter should be carefully done, since a small perturbation in such parameter leads to relevant quantitative changes. On the other hand, the estimation of a parameter with a rather small value for the sensitivity index does not require as much attention to estimate, because a small perturbation in that parameter leads to small changes. From Table 2, we conclude that the most sensitive parameters to the basic reproduction number R[1]c of the COVID-19 model (2.1) are β,p1,p2,η,σ,δ. In concrete, an increase of the value of β,p1,p2,σ will increase the basic reproduction number by 100%, 33.4%, 29.69%, 33.21% respectively and an increase of the value of η,δ will decrease R[1]c by 33.4% and 29.69% respectively. Sensitivity indices with respect to the parameter values as presented in Table 2 can be visualized from Figure 4.
4.2. Simulation results for model - 1
We have fitted the parameters β, p1, p2, p3, p4, μ, δ, σ, η, ρ1, ξ2 for the range of the values given in Table 1 and took ζ1, ζ2, ζ3, ξ1, ρ2, ν fixed as given in Table 1. We estimated those values for three different time intervals (1–30th day, 31–42nd day, 43–95th day). We have considered the initial values of the parameters as given in Table 2. We have chosen the time interval and end values of different compartments from the fittings over the previous time intervals. Here we have considered βSN(Is+p1Ia+p2E2+p3Q+p4J) as the daily reported case and the sum of this part over the desired time interval is considered as cumulative number of infected cases.
The optimization of parameters to describe the outbreak of COVID-19 in Germany was fitted by minimizing the Sum of Squared Errors (SSE), in such a way that the solutions obtained by the model approximate the reported cumulative number of infected cases. We applied three searches to minimize the SSE function: by using a gradient-based method first, followed by a step of minimization with a gradient-free method, again followed by a third step of gradient-based method. MATLAB based nonlinear least-square solver fmincon and patternsearch are used to fit simulated and observed daywise cumulative number of infected cases for three different time intervals. Detailed description of this method and its implementation can be found in [20,37,38,39].
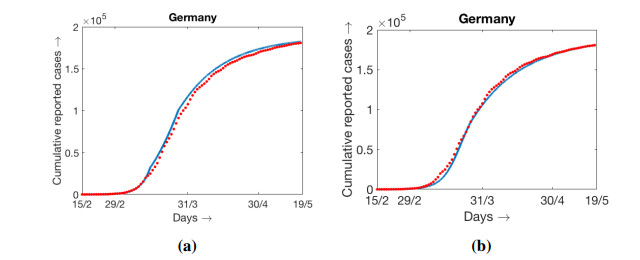
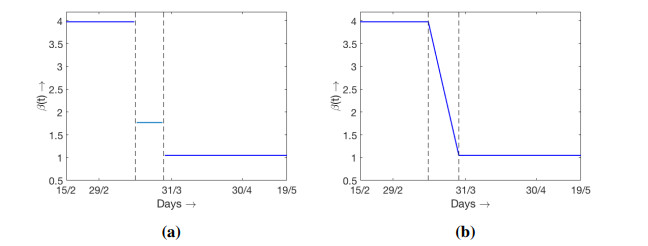
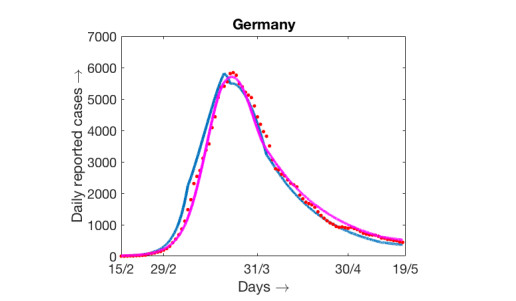
The model (2.1) is simulated with best fitted parameter values as mentioned above and the simulation result against the daily reported data and cumulative data are presented in Figure 5a. In this case the values of β are constants over three different range of days starting from the initial date of COVID-19 epidemic spread in Germany. For this figure some of the parameters are estimated as described above. Another simulation result is presented in Figure 5b where initially the value of β is constant for several days, and then monotone decreasing over a range of days and then remain constant for the rest of the period. The variation of β with respect to time is shown in Figure 6. Interestingly the data are fitted well in the case of time varying β where β(t) is continuous. For the simulation result presented in Figure 5b, the number of days for which β(t) remains constant and then starts decaying are adjusted in such a way that the outcome matches well with both the cumulative and daily data. Accordingly the slope of decaying β(t), the day up to which it decays and the lower value of β(t) are chosen. The numerical simulation and fitting with the data are carried out with the objective that the simulation result should be pretty close to the cumulative number of reported cases. We can claim that our attempt is successful as the cumulative number of reported cases for Germany on 15th May, 2020 obtained from the simulation is 180, 788 and the reported data (see [3]) shows it is equal to 182, 250. In this case the choice of β(t) is as shown in Figure 6a. Cumulative number of reported cases obtained from the simulation with the β(t) as shown in Figure 6b is equal to 180, 808. Both the simulated values are close to the reported data. Relative percentage error is less that 1% and is precisely equal to 0.79%.
Simulations and fitting are done with the reference to cumulative number of reported cases. However, the obtained results are also in good agreement with the daily data. As there is no uniformity of the daily reported cases, may be due to irregular reporting, we have used 7 days moving agerage as daily data instead of daily raw data. Simulation results with two different types of β(t) and 7 days moving average for daily reported data are shown in Figure 7. The estimated and other parameter values for the simulation with β(t) constant over three different time intervals are presented in Table 3 along with the sensitivity incides.
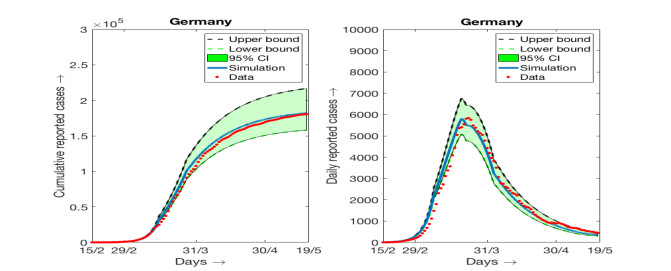
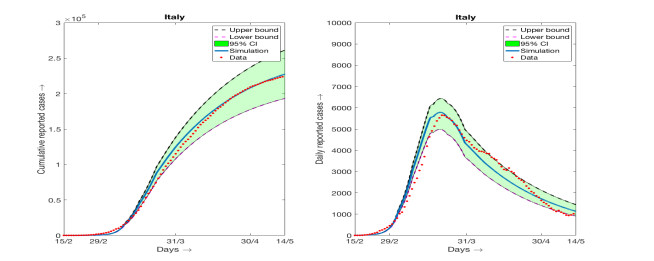
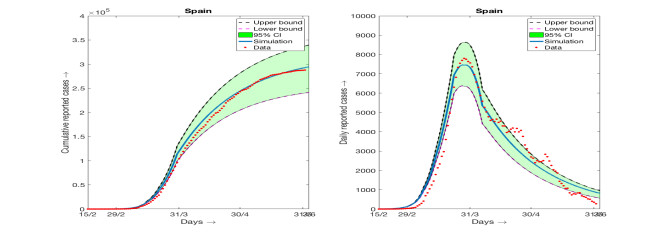
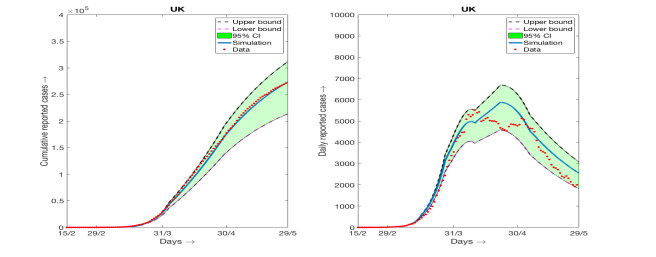
Consideration of changing β over different range of durations is an important observation of our present work. It is important to mention that the number of intervals over which we have to take different values of β is not always limited to three as in the case of Germany. The number of intervals over which we need to choose different values of β in order to match the simulation results with the data from different countries varies significantly. Of course, in all such simuations the values of β are decreasing with respect to the number of days. The simulations results for three more countries, namely Italy, UK and Spain, are provided in the appendix along with the result for Germany again where the 95% confidence intervals are shown. The parameter values used, the range of days over which β remains constant and estimated parameter values are also provided in the appendix.
4.3. Simulation results for model - 2
In order to understand the usefulness of the model (3.1), we now present some numerical simulation results. In Figure 5 we have presented the simulation results for the model (2.1) up to 19th of May. For the fitting with daily data, we have obtained three different values of β as β1=3.98, β2=1.77 and β3=1.05. The partial lockdown was started at Germany on 13th March, 2020 and then went into complete lock lockdown step by step. The process of unlocking at the essential sectors were started from 15th of May, 2020 and by that time the spreading speed of COVID-19 was much reduced. There are several restrictions like using mask, maitaining social distances and couple of other restrictions are in place in order to prevent the further spread of the disease. As of now the disease is not completely eradicated rather a minor number of cases are getting reported on a regular basis but the situation is seem to be under control.
We are interested to see the situation if all the citizens do not follow the suggested restrictions then what can happen afterward. For this purpose we now perform an interesting numerical simulations. We have simulation results for the model (2.1) as described at the previous subsection up to 19th of May. Now we start simulating the model (3.1) from the day immediately after 19th May and continue up to the end of November 2020. We assume that a fraction of population are not following the guideline and other fraction is following the safety and preventive norms appropriately. Let us define the ''coefficient of social interactions'' as S1/N and is denoted by K. If we assume that the 10% of the population is not following the norm then S1/N≡K=0.1 and hence S2/N=0.9=1−K. We assume that β11=β1, β22=β3 and β12=β21=(β1+β3)/2 where βj (j=1,2,3) are the values determined by fitting the data with the first model. Initial densities of E11 and E21 can be distributed with the same proportion to K and 1−K of the value of E1 on 19th May. Same procedure is adopted for other components except Q, J and R. For simplicity we also assume that the other parameters involved with the two group model are equal to the values of the corresponding parameter involved with the model (2.1) and is equal to the numerical values as mentioned at the last column of Table 3. For clearer understanding we can mention here some of the parameter values: pj1=p1=0.34 (j=1,2,3,4), σ1=σ2=σ=0.1 and so on.
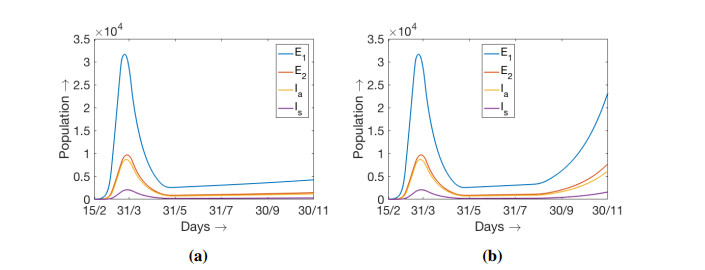
With above choice of initial values and parameter values now we simulate the model (3.1) up to the end of November, 2020. Two different simulations are performed and the simulation results are presented in Figure 8. For first simulation we have considered K=0.1 fixed up to the end of November 2020 (see Figure 8a). Second simulation is performed for K=0.1 upto the end of August and then K=0.15 during September to November 2020. In the figure we have ploted the consolidated compartments E1, E2, Ia and Is in order to avoid any confusion where we have obtained the values of these compartments by adding the values of the respective compartments in two group. To be specific, E1=E11+E12 and similary for other compartments. A relatively small increase of K lead to an essential acceleration in the disease progression.
With the parameter setup mentioned above one can calculate the controlled reproduction number R[2]c using the detailed formula given in Sub-section 3.1. Without any loss of generality we can make a crude assumption that N1N=K in order to find a value of R[2]c. Clearly the measure for N2N will be 1−K. With K=0.1, we can see slow growth in exposed and infected compartments as shown in Figure 8a as we find R[2]c=1.089. Now if we change the value of K to K=0.15 the revised measure of R[2]c becomes R[2]c=1.219. These results ensure the usefulness of our model along with the mathematical details and supportive numerical simulations.
5.
Model with relapse
Most of the important features and several issues related to COVID-19 remain unclear and immense scientific efforts are required to understand various important issues. As it is reported at many sources that some of the recovered individuals can become infected after a certain period of time as the recovery is seems to be temporary [40]. In this section we want to revisit the model (2.1) with the modification that a fraction of recovered population can return back to the susceptible class. We are mainly interested to calculate the final size of the infected compartment due to fact that the recovered individuals join the susceptible class after a certain number of days. In order to find a rough estimate of the size of the infected compartment and to obtain their estimate explicitly we can make a crude assumption. The assumption is that the disease related death rate is negligible. With these assumptions the model (2.1) with relapse of the disease can be written as follows
This model admits an endemic equilibrium point apart from the disease free equilibrium point. Let us denote the components of endemic equilibrium point as (ˉS,ˉE1,ˉE2,ˉIa,ˉIs,ˉQ,ˉJ,ˉR), then equating the right hand sides of (5.1c)–(5.1g) to zero, we find
Now equating the right hand sides of (5.1a) and (5.1b) to zero and then adding, we can find
Now we consider the following equation obtained from (5.1b),
Using the expressions for ˉE1, ˉE2, ˉIa, ˉQ, ˉJ in terms of ˉIs, we can write
and then using ˉS=N−ˉE1−ˉE2−ˉIa−ˉIs−ˉQ−ˉJ−ˉR and the relevant expressions we can find
where
In order to come up with a closed form of estimate for the endemic steady-state in case of relapse of the disease, we have assumed that the number of COVID-19 related death is negligible. In reality this is not true however it help us to have a rough estimate for the endemic steady-state dependning upon the value of ϵ. The parameter ϵ is related to the relapse rate as 1ϵ is the average number of days after which a recovered individual can have fresh infection. It is diffcult to derive explicit condition for which B1 is less than one in order to have a feasible values for the components of endemic equilibrium point in case of replase. In order to have a feasible values of ˉIs and other components we should have B1<1. Interestingly, with the parameter values mentioned at the last column of Table 3, we can calculate B1=1.12 where β=1.07 and note that R[1]c=0.7002. In order to have an admissible value of B1, if we consider β=1.2 and with other parameter values are same as in the third column of Table 3 we find B1=0.98 although R[1]c remains less than one. Finally assuming ϵ=0.01, that is recovered individuals can join the susceptible class after 100 days on an average, we find 1−B1B2≈6×10−5. So a rough idea about the estimated number of infected individuals at the endemic state will be 6×10−5×N where N is the total population of the country. The measure 1−B1B2 decreases with the decrease in ϵ as we can calculate 1−B1B2≈0.000031,0.000021 for ϵ=0.005,0.0033 respectively.
6.
Discussion
A wide range of mathematical approaches are adopted to develop viable mathematical model to understand the propagation of disease spread for COVID-19. The proposed mathematical models are not only diffierent in the context of basic assumptions behind the model formulation, rather incomplete or unclear understanding of disease spread for COVID-19 is another burden. In this paper we have proposed and analyzed an ordinary differential equation model to study the COVID-19 disease propagation which consists of fundamentally six compartments. The basic compartments are susceptible, exposed, infected, quarantined, hospitalised and recovered. It is evident that all the exposed individuals can not spread the disease and hence the exposed compartment is divided into two sub-compartments E1 and E2 where the individuals belonging to the second compartment can infect the healthy individuals. Once the incubation period is over, exposed individuals are becoming actively infected which means they can infect other individuals. In case of COVID-19 disease, every infected individuals are not developing appropriate symptoms and asymptomatic individuals can recovered from the disease without any serious medical intervention. The infected compartment is divided into two compartments, namely, asymptomatic infected and symptomatic infected.
There is a significant variation in the collection and reporting of infected data and no uniformity is maintained so far due to the rapid spread of the disease. From huge dataset, it is quite difficult to understand when an individual is identified as a COVID-patient but at which category they belong to (whether an individual belongs to E1, E2, Ia or Is compartment). Without any loss of generality we can assume that the individuals belonging to Ia and Is are tested to be positive. Once an individual is tested to be positive, he/she needs to follow the appropriate guideline of the country and hence they will be under isolation or quarantine assuming that they do not need any medical intervention through hospitalization. Hence we have assumed that the individuals from Ia compartment directly move to the recovered compartment and hospitalization is required for certain fraction of individuals belonging to sysmptomatic infected and quarantined compartments. COVID-19 related death is assumed for the Is and J compartments only. As the quarantined individuals are monitored on a regular basis and the healthcare service is supposed to be effective, hence every needy individuals can be moved to the hospital as per requirement. Based upon these assumptions we have considered two models here, epidemic model (2.1) and two group epidemic model (3.1). Interesting and significant contribution of our work is the consideration of time dependent rate of infection over various periods of time. This variation is adopted into the model in order to capture the effect of lockdown, social distancing etc. which play a crucial role to reduce the disease spread. The date of implementation of lockdown and the extent of lockdown varies from one country to another. Total lockdown is rarely implemented, rather most of the European countries went to step-by-step lockdown and sometimes they imposed complete lockdown at some states and/or provinces instead of total lockdown accross the country.
Parameter identification has a crucial importance in the epidemiological models. Some of the parameters, such as average durations of the incubation period and of hospitalization, can be evaluated from the clinical data. Some other parameters or their combinations are estimated by fitting the epidemiological data, in particular, the reported number of infected cases. However, even the combination of available methods of parameter identification does not usually allow a unique determination of all parameters. Different combinations of parameters can give a similar fitting accuracy. This means that the values of individual parameters cannot be uniquely determined. In spite of this uncertainty, the results of such modelling can have some predictive value because relative variation of parameters can be more important that their absolute values. As such, according to modelling, a slight increase of the coefficient of social interaction can lead to the second wave of epidemic, what is observed in Israel, Spain, France in July-September 2020. Therefore, we can evaluate necessary measures to stop this progression imposing stricter social distancing.
In this present work we have considered different durations of the period of time when the rate of infection varies in order to match the simulation results with the cumulative number of reported infected individuals for four countries. The magnitudes of β(t) over various intervals are obtained through fitting the simulated result with data but the choice of different periods remain in our hand. One can argue that the obtained results might change with a variant assumption but at the same time we can claim the appropriate nature of varying infectivity is not much clear yet. In order to show the effectiveness of our analysis, the fitting is done for the data from four different countries. It is important to mention here that the fitting with daily data seems to be not in good agreement (we can see significant variation around the fitted curve) but at the same time we need to keep in mind the irregularity of the reporting protocol. It is a matter of fact that several contries have changed their reporting mechanism time to time alongwith the change in the process of testing. A natural question may arise that we are considering constant β(t) over different time intervals, so what will be the outcome if we match the simulation result with the daily data or 7 days moving average? The answer is that either we need to change β(t) in daily basis which is not a good idea for fitting the simulation results of ordinary differential equation models with the data or we can observe a significant disagreement with the cumulative number of reported cases. For numerical simulation of the model (2.1) we have obtained three different values of β and we can calculate the controlled basic reproduction numbers accordingly. For the set of parameter values as mentioned in the three columns of Table 3, we can calculate the controlled reproduction numbers as R[1]c=2.9565,1.2847,0.7002. In Section 2 we have calculated the basic reproduction number R0, for model (2.1), apart from the controlled reproduction number R[1]c. If we calculate the basic reproduction number R[1]0 for model (2.1) with the parameter values given in the first column of Table 3, we find R[1]0=3.888 which is quite higher than R[1]c. It clearly indicates that the disease can propagate much faster in the absence of the control measures.
Epidemiological data on COVID-19 epidemic provide the number of registered infection cases on the bases of PCR tests, the number of hospitalized, recovered and dead (see, e.g., [3]). The number of positive tests depends on the total number of effectuated tests. However, if the organization of testing remains the same during some period of time, we can reasonably assume that the number of registered cases represents the total number of cases with some proportionality coefficient. In particular, if infected individuals apply to the medical care in the case of severe symptoms, subsequently tested and identified as positive cases, we can take into account that severe cases represent some given proportion of the total number of cases. Furthermore, since at the beginning of epidemic the number of susceptible individuals remains close to the total population, then the model is approximately linear, and the size of all sub-classes can be obtained by the same proportionality coefficient. On the other hand, some countries practiced a wide testing allowing a reliable estimate of the total number of infected cases. Overall, in spite of certain difficulties in the estimation of the number of infected individuals, we can reasonably assume that available epidemiological data faithfully reflects the real situation.
The proposed two group epidemic model can capture the situation when one group of people are following the social distancing norm, using face mask appropriately and other group is not obeying the safety norms. We must admit that the data and relevant information for the parameter values involved with the model (3.1) are not available yet. But the numerical simulation shows that if 10% or more of the population start violating the safety norms then there is a resonable chance that the disease can relapse. It is difficult to predict the size of the next epidemic but our model is capable to capture the realistic scenario. The number of daily reported cases started increasing at some countries like Iran, Spain etc. (see the data avialbale at [3]) compared to the number of reported infections on the days close the end of complete lockdown.
The coefficient of social interaction K introduced in Section 4.3 characterizes the intensity of the interaction between susceptible and infectious individuals, and it determines the rate of disease progression. The value K=0 corresponds to complete lockdown, and K=1 to the absence of lockdown. Partial measures of social distancing after the end of lockdown in a number of European countries restrain its value to K≈0.1 characterized by a slow growth of the number of infected individuals. Even a slight increase of this coefficient up to K=0.15, which can be expected in September due to the end of vacation season and the beginning of academic year, can lead to an important burst of epidemic.
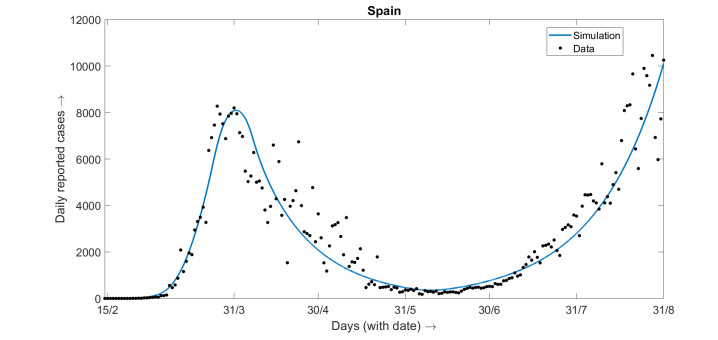
Currently available data for the unlocking period/post lockdown period indicates a significant increase in daily reported cases in Italy, full scale second wave of epidemic in Spain. There is not significant increase in number of case in Germany for which we have given much emphasis. Based upon the recently available data for Spain, we have fitted our 2nd model with the daily reported cases for Spain where second wave in epidemic has been noticed and got the best fitted value of K as 0.3 for which our model simulation has a good agreement with the real data for daily reported cases. The fitting with data in presented in Figure 9, all the parameter values are mentioned at the appendix. The choices of different parameters involved with the model (3.1) are the same as discussed in the previous section. The choices of βij are β11=β1, β22=β4 and β12=β21=(β1+β4)/2.
We have calculated the final size of the epidemic for both the models considered in this manuscript and obtained a rough estimate for the endemic steady-state analytically. The final size of the epidemic are calculated assuming β is constant however the numerical simulations are carried out for non-constant β and their variation with time is already explained clearly. Here we justify the analytical findings with supportive numerical results and argue for the effectivity of lockdown and imposition of several social restrictions. We mentioned above that the controlled reproduction number for Germany is R[1]c=2.9565 with β=β1=3.98. With this value of R[1]c, solving the equation (2.19) we find x=0.06303. With the current population size of Germany which is approximately equal to 82.9 million gives us final size as Sf=5,225,187. The 95% confidence interval for the final size is given by [4.7194×106,5.1958×106]. This result clearly indicates that a huge number of people could be infected if no such restriction was imposed. Similar calculations can be done for other compartments as well for two group model also. But we skip them here for the sake of brevity and will address this issue in our future endevour with appropriate estimates for the values of other parameters involved with the second model.
Control of the progression of COVID-19 epidemic goes beyond pure epidemiological questions. There are several societal aspects that can be important including the strategy chosen to handle the epidemic, and how people react on the measures of social distancing. In particular, at the first stage of epidemic, different measures were adopted according to the decisions of public authorities. Stricter measures of social distancing were introduced in China, while more liberal in Europe, especially, in Sweden. Tracing of infection chains with a large number of tests was applied in South Korea and in Germany. The results of these different strategies were also different, and they should be analyzed in more detail. The efficiency of lockdown introduced in many countries and post-lockdown evolution of the epidemiological situation depend on many factors. In particular, the application of the measure of social distancing can be influenced by some national traditional and historical features such as family meetings, festivities, religious meetings and holidays, and so on. Epidemic progression can also influence public opinion and mass media, which in their turn influence the intensity of social interactions [26,41].
Acknowledgments
The second author's (VV) work is supported by the Ministry of Science and Higher Education of the Russian Federation: agreement no. 075-03-2020-223/3 (FSSF-2020-0018).
Conflict of interest
The authors declare no conflicts of interest in this paper.
Appendix – A
Proof of Th. 1: By re-writing the system (2.1) we have
where
with
and so on. We note that
The inequalities mentioned above hold for any point belonging to the interior of R8+ or on the boundary hyper-planes. Solutions starting from a non-negative initial condition and non-negativity of the time derivaties imply that the system (2.1) is invariant in R8+.
Appendix – B
The matrices F1 and V1 are defined by
The matrices F1 and V1 are given by
Now, we can calculate,
F2 and V2 are given by
These two matrices F2 and V2 are given by
Explicit expressions for the matrices F3 and V3 are given below,
Appendix – C
Here we present the numerical simulation results and fitting with the data alongwith the 95% confidence intervals for Germany, Italy, Spain and UK. The data used for these fiiting are available at [3]. The model (2.1) is used to perform the simulations and fitting with data. Parameterization are also the same as we have explained in Table 1. Details of the parameter values and their estimates for Germany are provided in Table 3. Parameter values and and their estimates for rest of the countries are provided below. It is worthy to mention here that the values of β(t) are estimated over requisite number of intervals in order to have a good fit with the cumulative reported number of infected individuals. As a result the number of intervals over which β(t) is changing its constant magnitude varies from one country to another. In case of Germany, the number of intervals over which β(t) takes different constant values are three whereas the number of intervals for β(t) is five when we considered the data for Italy. Also the number days over which the simulations are carried out is not unique as the date implementation of lockdown and its partial withdrawal varies from one country to another. For all the countries, we have started the simulation from the date 15th of February, 2020 and continued approximately up to the end of strict lockdown restrictions.









 DownLoad:
DownLoad: